Full metadata record
| DC Field | Value | Language |
|---|---|---|
| dc.creator | Ferrer-Bonsoms, J.A. (Juan A.) | - |
| dc.creator | Cassol, I. (Ignacio) | - |
| dc.creator | Fernández-Acín, P. (Pablo) | - |
| dc.creator | Castilla, C. (Carlos) | - |
| dc.creator | Carazo-Melo, F.(Fernando) | - |
| dc.creator | Rubio, A. (Ángel) | - |
| dc.date.accessioned | 2023-06-02T09:26:18Z | - |
| dc.date.available | 2023-06-02T09:26:18Z | - |
| dc.date.issued | 2020 | - |
| dc.identifier.citation | Ferrer-Bonsoms, J.A. (Juan A.); Cassol, I. (Ignacio); Fernández-Acín, P. (Pablo); et al. "ISOGO: Functional annotation of protein-coding splice variants". Scientific Reports. 10 (1069), 2020, | es |
| dc.identifier.issn | 2045-2322 | - |
| dc.identifier.uri | https://hdl.handle.net/10171/66540 | - |
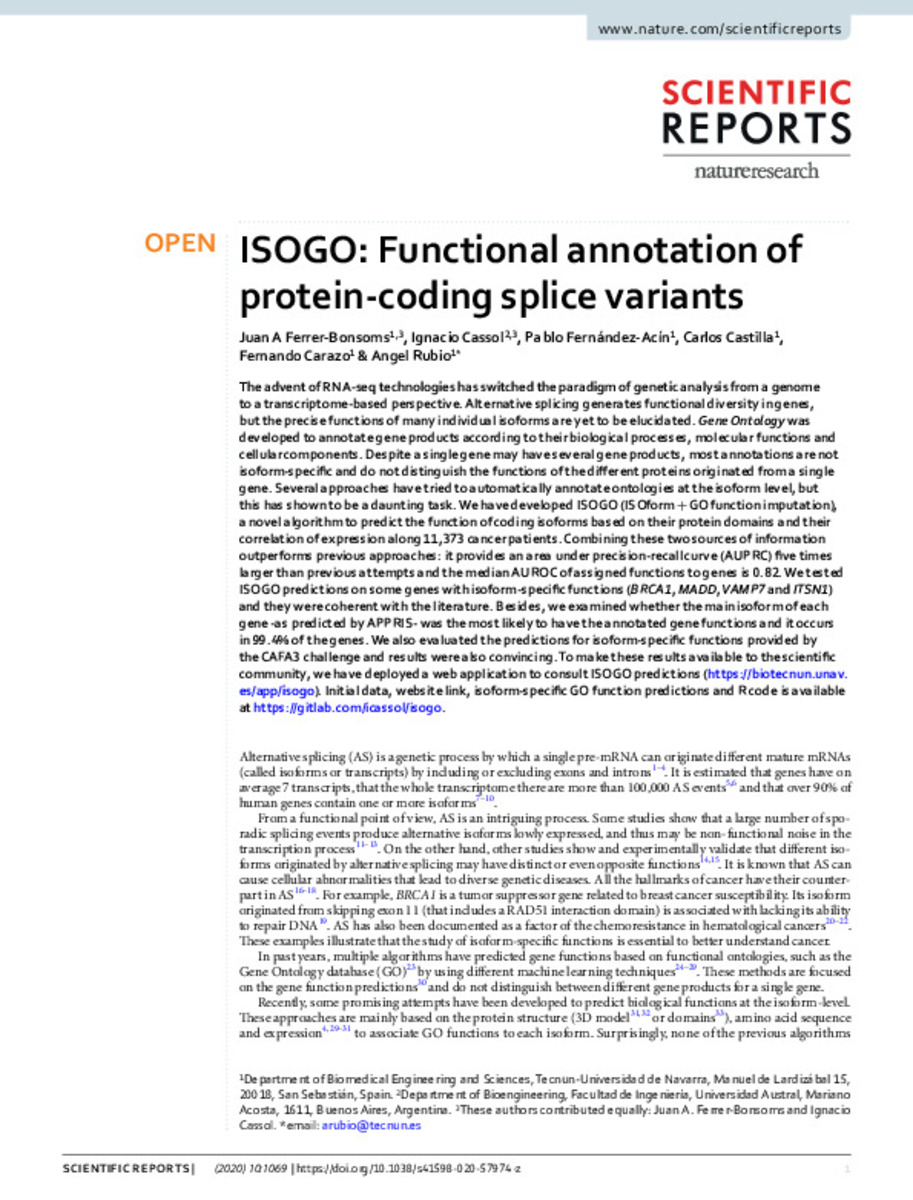
| dc.description.abstract | The advent of RNA-seq technologies has switched the paradigm of genetic analysis from a genome to a transcriptome-based perspective. Alternative splicing generates functional diversity in genes, but the precise functions of many individual isoforms are yet to be elucidated. Gene Ontology was developed to annotate gene products according to their biological processes, molecular functions and cellular components. Despite a single gene may have several gene products, most annotations are not isoform-specifc and do not distinguish the functions of the diferent proteins originated from a single gene. Several approaches have tried to automatically annotate ontologies at the isoform level, but this has shown to be a daunting task. We have developed ISOGO (ISOform+GO function imputation), a novel algorithm to predict the function of coding isoforms based on their protein domains and their correlation of expression along 11,373 cancer patients. Combining these two sources of information outperforms previous approaches: it provides an area under precision-recall curve (AUPRC) fve times larger than previous attempts and the median AUROC of assigned functions to genes is 0.82. We tested ISOGO predictions on some genes with isoform-specifc functions (BRCA1, MADD,VAMP7 and ITSN1) and they were coherent with the literature. Besides, we examined whether the main isoform of each gene -as predicted by APPRIS- was the most likely to have the annotated gene functions and it occurs in 99.4% of the genes. We also evaluated the predictions for isoform-specifc functions provided by the CAFA3 challenge and results were also convincing. To make these results available to the scientifc community, we have deployed a web application to consult ISOGO predictions (https://biotecnun.unav. es/app/isogo). Initial data, website link, isoform-specifc GO function predictions and R code is available at https://gitlab.com/icassol/isogo. | es_ES |
| dc.language.iso | eng | es_ES |
| dc.rights | info:eu-repo/semantics/openAccess | es_ES |
| dc.title | ISOGO: Functional annotation of protein-coding splice variants | es_ES |
| dc.type | info:eu-repo/semantics/article | es_ES |
| dc.description.note | This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. Te images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/. | es_ES |
| dc.editorial.note | Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional afliations. | es_ES |
| dc.identifier.doi | 10.1038/s41598-020-57974-z | - |
| dadun.citation.number | 1069 | es_ES |
| dadun.citation.publicationName | Scientific Reports | es_ES |
| dadun.citation.volume | 10 | es_ES |
Files in This Item:
Statistics and impact
Items in Dadun are protected by copyright, with all rights reserved, unless otherwise indicated.